tripso
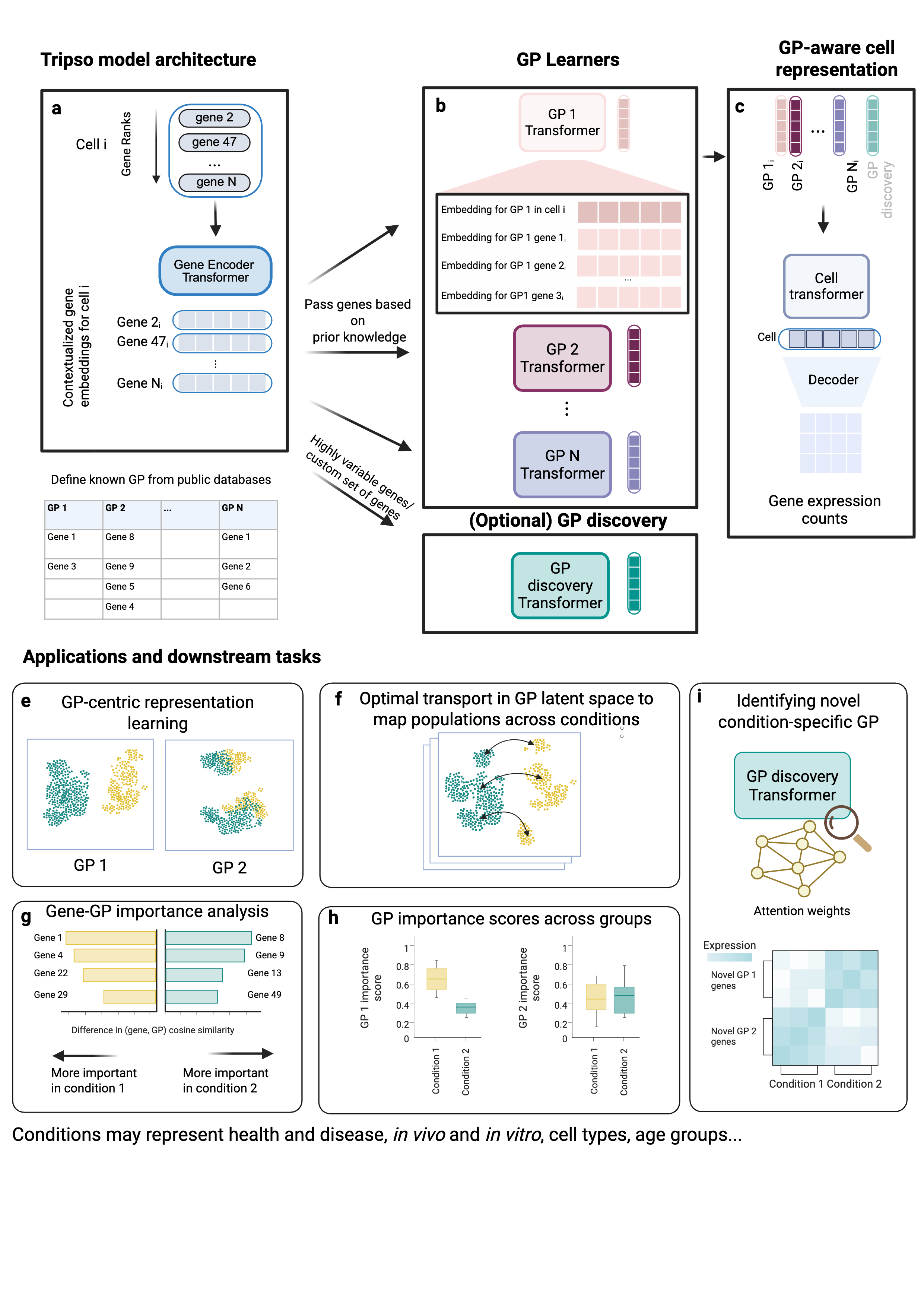
Welcome to the Tripso documentation! Tripso (Transformers for learning Representations of Interpretable gene Programs in Single-cell transcriptOmics), is a self-supervised framework that decomposes cellular states into multiple GP embeddings. Tripso learns contextualised gene embeddings within each GP, quantifies gene-level contributions to each GP, as well as this influence of each GP on global cell identity.
You will find links to tutorial scripts and installation instructions below.
Getting Started
Tutorials
- Data Preparation
- Gene Program Database Creation
- Cell Tokenization
- Train Base and Global Models
- Model Evaluation and Gene Program Importance Analysis
- Visualizing Gene Program Embeddings
- Calculating Gene Program (GP) Importance Scores
- Generating GP Gene Cosine Similarity
- Analyzing Gene-to-GP Cosine Similarity
- Extracting novel gene programs
- Visualizing Novel Gene Programs Discovered by GPFinder
Contributing